- Home
- Applications
- Measure Cell Migration and Chemotaxis in Real Time
- Incucyte® Chemotaxis Assay
- Chemotaxis Application Note
Application Note
Quantification of Cell Migration Using the IncuCyte® Chemotaxis Assay
Introduction
Cell migration is an essential component of most biological processes, including: immune responses, blood vessel formation, tumor metastasis, and wound healing [1]. Effective in vitro models of directed cell migration, or chemotaxis, are necessary for both basic research and drug discovery [2]. This application note describes a new methodology for studying chemotactic cell migration using the IncuCyte® ZOOM Live-Cell Imaging system. This novel approach enables real-time visualization and automated analysis of chemotactic migration in a 96-well format within a tissue culture incubator.
By incorporating a made-for-purpose, optically clear membrane into a transmembrane geometry, the IncuCyte® chemotaxis assay facilitates the label-free visualization and quantitation of directed cell migration. The design of the IncuCyte® ClearView 96-Well Cell Migration Plate ensures a stable gradient over 72 hours and requires only 1,000–5,000 cells per well. In contrast to traditional Boydenchamber methodologies, cells in the IncuCyte® chemotaxis assay are required to migrate on a biologically relevant surface toward the chemoattractant. The result is 96 wells of reproducible kinetic data supported by images and movies, requiring very few cells and minimal hands-on time.

Figure 1. Assay Workflow
- Most cell types tested require a coating of the membrane surface to facilitate chemotactic migration.
- Subconfluent cells are harvested, counted and plated in the upper chamber at densities ranging from 1,000 (adherent cells)–5,000 (non-adherent cells; T cells and neutrophils) cells/well.
- Compounds, if required, are added to the upper chamber and mixed.
- The plate is incubated briefly at room temperature to allow cells to settle.
- Chemoattractant is added to the reservoir, the upper chamber is placed into the reservoir, and the assay is initiated in the IncuCyte® ZOOM instrument.
Data Quantification and Analysis
Whole-well images of cells on both the bottom and the top of the ClearView plate membrane are captured at user-defined intervals. All images are processed using automated algorithms to quantify cell area on each side of the membrane (Figure 2). Directed cell migration can be reported as either an increase in area on the bottom side of the membrane for adherent cells (Figure 3), or a decrease in area on the top side of the membrane for non-adherent cells that migrate down the pore and fall off of the membrane (Figure 4).
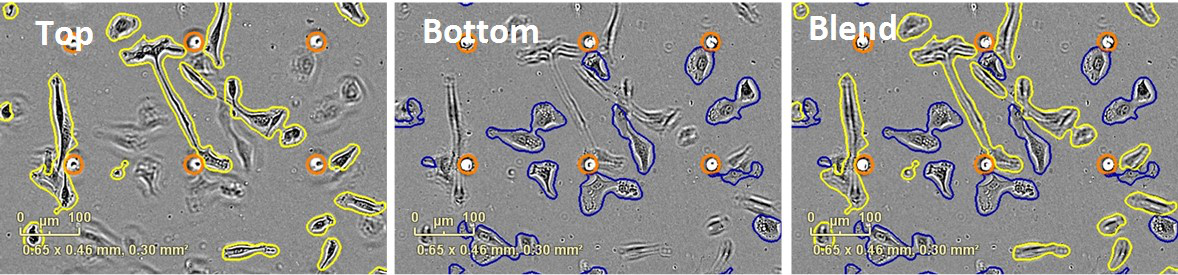
Figure 2. Chemotaxis quantification. HT-1080 fibrosarcoma cells were plated in the top chamber of the IncuCyte® ClearView 96-Well Cell Migration Plate at a density of 500 cells/well. 10% FBS was added to the bottom chamber as a chemoattractant. Images represent the top and bottom side of the membrane at the 36-hour time point. Automated image processing separates cells located on the top (outlined in yellow) and the bottom (outlined in blue) surface of the membrane. Pores are outlined in orange. Images are processed as they are acquired, and data can be plotted in real time.
Cancer Cell Chemotaxis (Adherent Cells)
In a model of cancer cell migration, we investigated the pharmacological eff ect of traditional signaling pathway inhibitors on the directed migration of HT-1080 fibrosarcoma cells toward fetal bovine serum (FBS) (Figure 3). We observed concentration dependent inhibition of directed cell migration with each of the three inhibitors tested: KU0063794 (Akt pathway), U0126 (MEK/ERK MAPK pathway), and Wortmannin (PI3K pathway).

Figure 3. Inhibition of HT-1080 cancer cell migration. This 96-well microplate graph illustrates the kinetic measurement of HT-1080 cell migration toward 10% FBS in each well of the IncuCyte® ClearView 96-Well Cell Migration Plate. Plotted in each well is the cell area on the bottom of the membrane (y-axis) over the course of a 48-hour assay (x-axis). U0126, KU0063974 or Wortmannin were added to 500 HT-1080 cells in the upper chamber and incubated at 37°C for approximately 30 minutes prior to exposing the cells to chemoattractant in the lower chamber. The concentration of inhibitors decreases from the top of the plate to the bottom. Positive (with chemoattractant) and negative (without chemoattractant) controls are located in the last row. These data were collected over a 48-hour period at 1-hour intervals.
Immune Cell Chemotaxis (Non-Adherent Cells)
In a model of immune cell chemotaxis, we investigated the response of CD3/CD28 activated T cells toward two chemoattractants: CXCL11 and CXCL12 (aka SDF-1α), ligands for CXCR3 and CXCR4, respectively (Figure 4) [3]. By measuring the loss of cell area on the top of the membrane, we show that activated T cells migrate toward both CXCL11 and CXCL12. Furthermore, we show that the selective CXCR4 antagonist, AMD3100, inhibits chemotaxis toward CXCL12 (IC50 = 279 nM), with no effect on CXCL11-mediated chemotaxis. This experiment was completed on an ICAM-1–coated surface. Interestingly, successful measurements of T cell chemotaxis were also made on fibronectin and Matrigel/FBS–coated surfaces, while T cells did not migrate on an uncoated surface. This suggests that interactions between integrins and/or receptors on the cell surface and the substrate play a fundamental role in T cell chemotaxis in this assay. This was not the case when tested in a traditional Boyden chamber, where the surface coating was not required (data not shown).
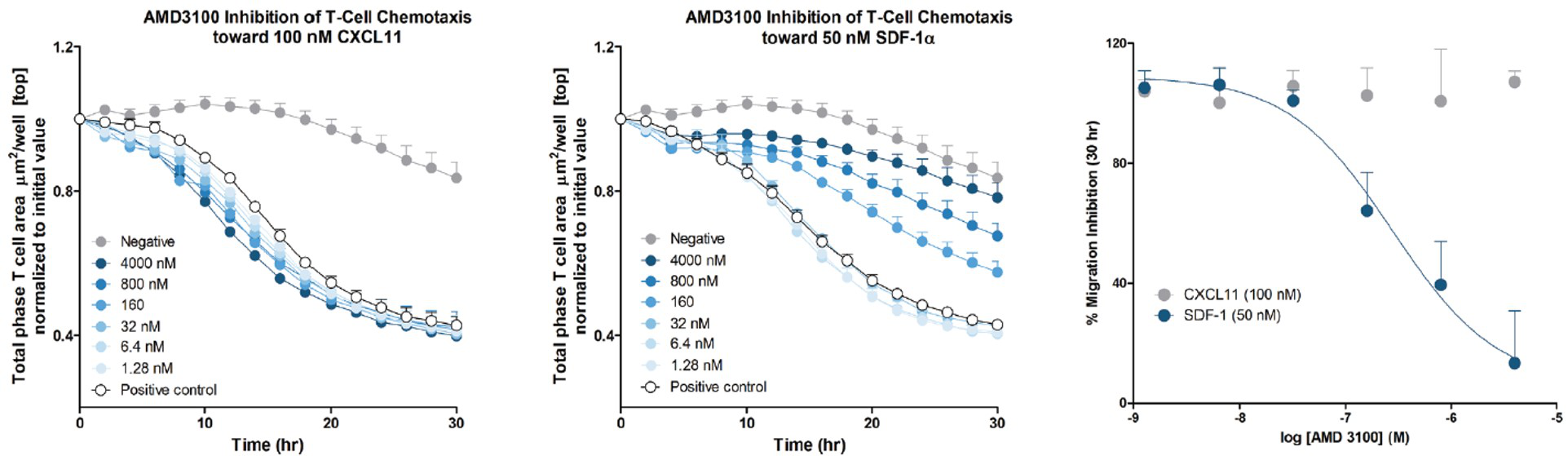
Figure 4. T cell chemotaxis toward CXCL11 and CXCL12 (SDF-1α). T-cells were plated at a density of 5,000 cells per well in the upper chamber of an IncuCyte® ClearView 96-Well Cell Migration Plate coated with ICAM-1. AMD3100 was added to the cells at indicated concentrations and incubated at room temperature for approximately 1 hour prior to exposing the cells to chemoattractant gradient. CXC11 or SDF-1α was added to the reservoir plate at 100 nM or 50 nM, respectively, based on EC50 values obtained in agonist curve experiments (data not shown). Analysis of pharmacological response was performed at t=30 hours. Data were collected over a 30-hour period at 2-hour intervals. Each data point represents mean ± SEM, n=3.
Intra- and Inter-Plate Reproducibility Measurements
To assess assay reproducibility and precision of the IncuCyte® chemotaxis assay, we performed a series of four independent experiments using Jurkat (non-adherent) T cells (Figure 5). Cells were plated at a density of 5,000 cells per well. Two-fold dilutions of CXCL12 were made across columns, and measurements of total phase object area normalized to the initial value were plotted over a 30 hour time course. Highly reproducible well-to-well kinetic measurements, with an average intra-assay CV of 6.3%, were observed. Similar results were obtained when measuring the directed migration of HT-1080 cells toward FBS and neutrophils toward IL-8, C5a and fMLP (data not shown).
Figure 5. Precision and reproducibility of directed migration of Jurkat T cells
toward CXCL12. Jurkat cells were plated at a density of 5,000 cells per well in the upper chamber of the ClearView cell migration plate. Two-fold dilutions of SDF-1α (n=8 per concentration) were added to the reservoir.
Stable Gradient
A long-term, stable gradient is required to support chemotaxis over longer periods of time, and to increase cell participation rates. In an experiment designed to directly compare the stability of a chemoattractant gradient in the IncuCyte® ClearView 96-Well Cell Migration Plate and the Corning® Transwell® Plate, diffusion of a 10 kDa dextran fluorescently labeled with Alexa Fluor® 594 was monitored over a 72 hour time course in each consumable. As shown in Figure 6A, more than 50% of the gradient dissipates in the Corning® Transwell® Plate within the first four hours. In contrast, >80% of the gradient remains intact in the IncuCyte® ClearView plate at 72 hours. In a biological test of gradient stability, the HT-1080 cells migrated directionally toward FBS with equal rates in gradients that had been pre-established for 24, 48, and 72 hours, indicating a stable gradient across the membrane (Figure 6B). These rates were slightly faster than those for cells migrating toward a gradient that had not been pre-established. This is likely a result of increased cell health when serum-starved cells are plated in established gradients.

Figure 6. Evaluation of ClearView chemotaxis assay gradient. (A) 10 kDa dextran, labeled with Alexa Fluor® 594, was added to the ClearView reservoir plate at a concentration of 10 μM to establish gradients over 72, 48, 24 and t=0 hours. Measurements of diffusion were made by sampling the ClearView insert wells and measuring fluorescent intensity on a microplate reader. Each data point represents mean ± SEM, n=3. (B). An FBS gradient was established over 72, 48, 24 and t=0 hours in separate wells. Serum starved NucLight Red HT-1080 cells were added at a density of 500 cells per insert well to the established gradients and FBS-induced chemotactic response was measured. Each data point represents mean ± SEM, n=3.
Conclusions
We have demonstrated that the IncuCyte® chemotaxis assay is a quantitative and reproducible approach for measuring chemotaxis in both adherent and non-adherent cell types. This assay format allows for the kinetic detection of cell migration toward chemotactic gradients on a physiologically relevant substrate, with movies and images that support quantitative measurements and provide associated morphological and phenotypic insights. Key features demonstrated in the IncuCyte® chemotaxis assay are:
- Visually monitor every cell in your experiment, assess morphology and gain phenotypic insight from images and movies
- Measure label-free or labeled cell migration without fixing, staining or cell scraping steps
- Set up and walk away – fully automated image-based quantification
- Highly reproducible 96-well approach suitable for profiling and screening
- Investigate cell migration on biologically relevant surfaces
- Sustained and stable gradient over 72 hours
Approach and Methods
Cell Culture
All cell culture reagents were obtained from Life Technologies unless otherwise noted. Cells (HT-1080 and Jurkat) were grown to subconfluence in either 25 cm2 or 75 cm2 tissue culture treated flasks prior to initiation of assays. HT-1080 cells were cultured in DMEM supplemented with 10% FBS, non-essential amino acids, sodium pyruvate, and 2 mM GlutaMAX™ supplement. Jurkat cells were cultured in RPMI-1640 (ATCC #30-2001) + 10% FBS.
T Cell Isolation
T cells were isolated from human whole blood using Lympholyte®-H cell separation medium (CedarLane #CL5010). Monocytes were removed from culture after adherence to tissue culture treated plastic for 1 to 6 hours. Remaining lymphocytes were transferred to a T75 culture flask and incubated with for 7 days with 25 ng/mL IL-2 (R&D Systems #202-IL) and CD3/CD28 Dynabeads® (Life Technologies #11131D) at a ratio of 1:1.
References
1 Ridley AJ et al. Cell Migration: integrating signals from front to back. Science 2003. 302, 1704-1709.
2. Hulkower K and Herber R Cell migration and invasion assays as tools for drug discovery. Pharmaceutics 2001, 3: 107-124.
3. Kawaguchi A, Orba Y et al: Inhibition of the SDF-1α-CXCR4 axis by the CXCR4 antagonist AMD3100 suppreses the migration of cultured cells from ATL patients and murine lymphoblastoid cells from HTLV-U Tax transgenic mice. Blood 2009, 114: 2961-2968.
