- Home
- News & Events
- News
- IncuCyte® e-Newsletter
- Spring 2016

In This Issue
Featured Application
 |
Cell Health & Viability Assays |
Just Released
 |
IncuCyte™ NucLight™ BacMam 3.0 Reagents IncuCyte™ Cytotox Reagents for counting dead cells |
Featured Products
 |
IncuCyte™ NucLight™ Reagents for nuclear labeling IncuCyte™ CytoLight™ Reagents for cytoplasmic labeling |
Publication Updates
In every issue, learn how researchers around the world are using IncuCyte™ systems across a range of applications and research areas. Read publications now
IncuCyte™ Tips and Tricks
Ask an Expert
Find answers to questions submitted by readers just like you. See latest answers
Discounts & Promotions
Get updates on our latest discounts and promotions. See our newest discounts and promotions
Upcoming events
Find out what's happening near you. See latest events
FEATURED APPLICATIONS
Cell health & viability assays
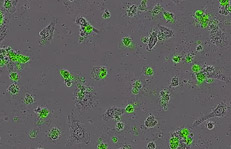
What are cell health & viability assays?
Cell health and viability assays enable the measurements of cell health, which are essential when studying the effects of drugs, culture conditions or genetic modifications on cell growth or viability. These assays also provide crucial means for ranking compounds in drug discovery screens and enable investigation into the cellular changes that underlie disease pathologies.
What is our solution?
Many existing methods for assessing cell health and viability rely on indirect, end-point measurements that are subject to artefacts and cannot be readily verified by morphology changes. IncuCyte™ Cell health and viability assays provide direct, image-based cell health measurements in real time within a standard cell culture incubator.
What it offers
- Automated label-free (confluence) measurements of cell growth and growth inhibition.
- Visualization and validation of changes in cell health in adherent or suspension cells, in co-cultures and 3D models. Ideal for cancer, immune and neuronal cell studies.
- Morphological and functional information from every well via time courses and concentration depend responses
Learn more about cell health & viability assays
Related applications
- Proliferation: Measure label-free cell growth or growth inhibition and count living cells in real time. Learn more
- Apoptosis: Detect apoptosis in living cells and in real time using simple mix-and-read protocols Learn more
- Cytotoxicity: Real-time measurements of cell viability using simple mix-and-read protocols suitable for screening Learn more
- 3D biology: Monitor and quantify 3D- spheroid formation, growth and shrinkage in real time Learn more
JUST RELEASED
IncuCyte™ NucLight™ BacMam 3.0 Reagents
What it is
The NucLight™ BacMam 3.0 reagents are used for non-perturbing, nuclear labeling of living mammalian cells. Using a simple mix-and-read protocol, these reagents enable real-time cell counting within hours of infection, thus allowing calculations of doubling times and direct measures of cellular proliferation. The reagents are optimized for the IncuCyte® system - you don’t need to worry about compatibility.
What it offers
- Rapid, non-perturbing, nuclear labeling of a variety of mammalian cell types
- Real-time quantitation of cell proliferation
- Experimentally and environmentally safe
- 96-well throughput for accurately counting cells in a population
Ordering info
- IncuCyte™ NucLight™ Red BacMam 3.0 Reagent for nuclear labeling (cat. no. 4621)
- IncuCyte™ NucLight™ Green BacMam 3.0 Reagent for nuclear labeling (cat. no. 4622)
IncuCyte™ Cytotox Reagents for counting dead cells
What it is
The IncuCyte™ Cytotoxicity Assays measure cell death based on cell membrane integrity in real time, all within your incubator. Using a simple mix-and-read protocol, these reagents enable real-time quantification of cell death. The reagents are optimized for the IncuCyte® system – you don’t need to worry about compatibility.
What it offers
- Visualize and quantify cytotoxicity using time-lapse imaging
- Automatic analysis of the time course of cell death within your incubator
- Simple mix-and-read 96/384-well protocols - no washing, no fixing, no lifting
Ordering info
- IncuCyte™ Cytotox Red Reagent for counting dead cells (cat. no. 4632)
- IncuCyte™ Cytotox Green Reagent for counting dead cells (cat. no. 4633)
FEATURED PRODUCTS
IncuCyte™ NucLight™ Reagents for nuclear labeling
What it is
IncuCyte™ NucLight™ Reagents enable efficient nuclear labeling and counting of living cells without perturbing cell function or morphology. They are a unique range of BacMam and Lentiviral-based labeling reagents with simple transduction protocols that enable expression of a nuclear-restricted green (GFP) or red (mKate2) fluorescent protein in your choice of primary, immortalized, dividing, or non-dividing cells.
What it offers
- Count living cells in real time over days – calculate true doubling times and growth rates
- Simple labeling protocols – compatible with your choice of cells
- Idea for co-cultures and multiplexing with IncuCyte™ apoptosis or cytotoxicity assays
Learn more about IncuCyte™ NucLight™ reagents
IncuCyte™ CytoLight™ Reagents for cytoplasmic labeling
What it is
IncuCyte™ CytoLight™ Reagents enable efficient cytoplasmic labeling of living cells and monitoring cell-cell interactions without perturbing function or morphology. They are a unique range of lentiviral-based labeling reagents with simple transduction protocols that measure cell health in 3D spheroids and enable expression of a cytoplasmically soluble green (GFP) or red (mKate2) fluorescent protein in your choice of primary, immortalized, dividing, or non-dividing cells.
What it offers
- Monitor the assembly of multicellular structures in real time over days or weeks
- Simple labeling protocols – compatible with your choice of cells
- Ideal for co-cultures and multiplexing with IncuCyte™ apoptosis or cytotoxicity assays
Publication Updates from the IncuCyte™ Community
Over 700 peer-reviewed publications* in research areas including:
| Research area | Publications |
|---|---|
| Oncology | 504 |
| Cardiovascular | 27 |
| Developmental biology | 35 |
| Immunology | 29 |
| Infectious diseases | 21 |
| Inflammation | 9 |
| Metabolism | 35 |
| Neuroscience | 35 |
| Regenerative medicine | 53 |
| Toxicology | 16 |
*Publications may appear in more than one research category.
Recent publication highlights
1. Park, S.H. et al. Vascular Protective Effect of an Ethanol Extract of Camellia japonica Fruit: Endothelium-Dependent Relaxation of Coronary Artery and Reduction of Smooth Muscle Cell Migration. Oxid. Med. Cell. Longev. 2016, 1–9 (2016).
2. Neri, S. et al. Cancer cell invasion driven by extracellular matrix remodeling is dependent on the properties of cancer-associated fibroblasts. J. Cancer Res. Clin. Oncol. 142, 437–446 (2016).
3. Karnezis, A. N. et al. Dual loss of the SWI/SNF complex ATPases SMARCA4/BRG1 and SMARCA2/BRM is highly sensitive and specific for small cell carcinoma of the ovary, hypercalcaemic type. J. Pathol. 238, 389–400 (2016).
4. Jamshidi, F. et al. The genomic landscape of epithelioid sarcoma cell lines and tumours. J. Pathol. 238, 63–73 (2016).
How can I improve my assay performance?
Experimental data is influenced by many factors including initial growth conditions, cell counting, and dissociation methods. In order to improve cell performance we recommend the following techniques:
1) Prior to assay initiation maintain cell culture at or below 80% confluence.
2) Ensure that you have accurate cell counts by thoroughly mixing your cells prior to sampling.
3) Limit the time the cells are exposed to enzymatic dissociation.
Q: How do I compare the growth of the same cells while they are plated in two separate flasks at the same starting density?
A: The growth curve of each flask must be generated separately, creating two separate graphs. For their direct growth comparison you can combine both graphs into one by selecting “Graph” on Graph Option in the Drag and Drop box at the bottom of the graph window. Simply right-click on, and hold the graph to be dragged, and then use the mouse to drag it onto the target graph.
Release of the mouse button will cause the graph to be dropped. The new graph will still contain the title from the original, Growth Curve for Vessel 1- however, it will now contain two traces labeled as Vessel 1 and Vessel 2.
Watch the video:
Have a technical support question?
Refer A Friend
Our IncuCyte™ Referral Program is a program that rewards you when someone you refer to us, purchases an IncuCyte™ system.
Come and visit us at these 2016 events.
March
- March 17-19: The 15th Congress of the Japanese Society for Regenerative Medicine, Osaka, Japan
- March 21-23: 3rd Immunotherapy of Cancer Conference (ITOC3), Munich, Germany
April
- April 17-20: American Association for Cancer Research (AACR), New Orleans, Louisiana, USA
May
- May 3-4: Lab Automation, Formulation & Phenotypic Screening, Elrig, Rennes, France
- May 10-12: 14th Annual Cancer Immunotherapy (CIMT), Mainz, Germany
- May 14-16: The American Association of Immunologists (AAI), Seattle, Washington, USA
- May 30-June 1: ACTC – Advances in Cell and Tissue Culture, Barcelona, Spain
- May 31: IncuCyte User Group Meeting, Frankfurt, Germany
June
- June 15-17: The 68th Annual Meeting of the Japan Society for Cell Biology, Kyoto, Japan